The trials and tribulations of identifying rhino parents
17/02/2020 in Conservation
The RZSS WildGenes Lab has been working in collaboration with the Institute of Biodiversity Animal Health and Comparative Medicine at the University of Glasgow on a project that aims to inform the conservation of Botswanan Rhinos. Tarid Purisotayo, a PhD student working on the project, is here to share the story so far.

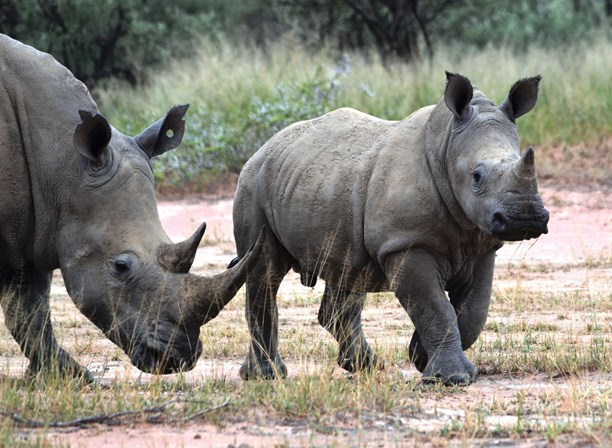
White rhinos are the larger of the two African species [credit: Nicholas Jonsson]
Initially, the primary aim of the project was to construct pedigrees for three southern white rhino populations in the Republic of Botswana, which essentially means piecing together a rhinos family tree! However, as it can often be in the world of genetics - things were more challenging than planned.
A brief history of rhino conservation
To explain what we have been working on, let me first run through a brief history of rhino conservation: During the mid to late nineteenth century, southern white rhinos were hunted until fewer than 100 individuals survived in one last stronghold in the Hluhluwe–Umfolozi Park in Kwazulu-Natal, South Africa. Luckily, this area became a national park in 1895 (the first in Africa) and with intensive protection, the population rapidly increased.
By the 1960s there were so many rhinos that Hluhluwe–Umfolozi was reaching what is known as its ‘carrying capacity’: the rhino population was in danger of exceeding the available food and resources. By this time however catching and anaesthetic techniques had developed so significantly that translocations into other suitable habitats were a viable option. This proved to be an efficient strategy, the southern white rhino reached a milestone of 20,000 individuals in 2012, and more translocations are now required.
Keeping track of rhino populations today
The majority of these rhino habitats are kept in fenced and intensively protected areas, and it was from the southern white rhinos of Hluhluwe–Umfolozi that most, if not all, re-introduced populations began. Because of this, it has been important to transfer unrelated rhinos between populations to avoid inbreeding. The challenge is keeping track of which rhinos are closely related and which rhinos are unrelated, up until now, such information has been based purely on ranger observations.

Fieldwork on Southern White Rhinos in Botswana [credit: Nicholas Jonsson]
Fortunately, scientists have other tools to trace the pedigrees, and this is where our research project steps in. We are using DNA analysis to put together family trees, or in technical speak, using DNA markers to resolve parentage. A DNA marker is a gene or DNA fragment that is always found in the same location on the genome. It is unique to a species or individual so can be used to identify one from another. Often, just ten individual DNA markers are enough to work out who is related to whom, however, things are complicated for the southern white rhino since all the descendants are very closely related.
Unfortunately, only twenty-three DNA markers were available for the species and our research project has confirmed that there is not enough difference between them to construct a complete family tree, even when combined with observational data from the rangers (find out more about this here).
The seach for genetic fingerprints
We needed to locate more markers to find unique genetic fingerprints, and we were lucky to obtain samples from almost all southern white rhinos within private-managed populations in the Republic of Botswana (thanks to Erik Verreynne our research partner from Agric Vet Consultants).
However, developing new DNA markers is not simple. We trialled two different methods, firstly, reduced representation sequencing techniques, and secondly, targeted capture sequencing. The former group of techniques does not require prior genetic knowledge but does require good quality DNA. The latter techniques, on the contrary, perform well with low quality DNA but requires prior genetic knowledge of the species. Unfortunately for the southern white rhino, we could not use either of them alone.

Hot field environments are not the ideal conditions to store samples! [credit: Nicholas Jonsson]
The first technique, reduced representation sequencing, has been widely used in population genetic analyses for conservation because it is a cost-effective way to analyse large numbers of individuals. But, our samples were collected from populations living in natural habitats where there are few facilities able to store them. The quality of DNA samples has been one major issue for our project as more than half of the samples were damaged, so we could not use this technique alone. Our research project would have to pave a new path.
Fortunately, European zoological parks under the European Endangered Species Programme and the Science and Advice for Scottish Agriculture (SASA) generously provided us with additional samples containing good quality DNA that we could use to develop an alternative approach. After we had obtained some successful sequencing from reduced representation techniques, we were able to identify new DNA markers. From 23 traditional DNA markers, we now had an incredible 8,000 candidate markers!
A hopeful future for the southern white rhino
So now with the information of these thousands of DNA markers, we should be able to work out the parents for most offspring. With this information, authorities will be able to decide which rhino to move where to ensure decent genetic diversity in each population. We hope this set of new DNA markers will benefit many research studies and the conservation of rhinos into the future.
Until next time,
Tarid


This work has been made possible by a partnership between the University of Glasgow and the WildGenes Lab, this work has been funded by a PhD grant from the Royal Thai government. We thank the Department of Wildlife and National Parks of Botswana for providing research permits, and the Vet and Agric Consultants in Botswana and the Science and Advice for Scottish Agriculture for providing samples.
We would like to thank the players of People’s Postcode Lottery for their generous support of the RZSS WildGenes lab.



























Follow EZ