Meet the conservation and veterinary team
Conservation team

HEAD OF CONSERVATION AND SCIENCE PROGRAMMES
Dr Helen Senn
Helen manages the RZSS conservation department. She leads a diverse team of field scientists, geneticists, animal managers and conservation planners who are enacting the RZSS pledge to reverse the decline of 50 species by 2030. She is the project lead for Saving Wildcats.
Contact: hsenn@rzss.org.uk

CONSERVATION PROGRAMME MANAGER
Dr Helen Taylor
Helen leads the conservation field team’s work on a wide variety of species, from tiny pine hoverflies to elusive giant armadillos. She has experience in managing conservation breeding and translocation programmes and is an IUCN CPSG-accredited Conservation Planning facilitator.
Contact htaylor@rzss.org.uk

CONSERVATION PROGRAMME MANAGER (RZSS WildGenes)
Dr Alex Ball
Alex manages the RZSS WildGenes team, the only zoo-based conservation genetics laboratory in the UK. Using genetics to inform conservation management, Alex’s team consists of laboratory technicians, scientists and biobank staff. The team works on 15-20 species annually in its efforts to support the conservation department’s wider mission.
Contact: aball@rzss.org.uk

SENIOR LABORATORY TECHNICIAN (RZSS WildGenes)
Elizabeth Heap
Liz is the senior laboratory technician at RZSS WildGenes’ conservation genetics laboratory based in Edinburgh Zoo. The lab delivers genetics and genomics data to the team of onsite scientists, supporting the conversation projects at RZSS. Liz also works on international capacity building projects and supports the WildGenes Biobank.
Contact: eheap@rzss.org.uk

RESEARCH SCIENTIST (RZSS WildGenes)
Dr Heather Ritchie-Parker
Heather is a scientist in the RZSS WildGenes team, analysing genetic and genomic date produced by the lab team to inform conservation management decisions. Heather is focusing currently on our penguin and native invertebrate projects.
Contact: hritchieparker@rzss.org.uk

CONSERVATION PROJECT OFFICER
Kasia Ruta
Kasia is on the conservation field team. Currently her main focus is on supporting conservation efforts for the charismatic Pallas’s cat and other elusive small cats across Central Asia, as well as delivering a conservation translocation programme for the little-known pond mud snail in Scotland.
Contact: kruta@rzss.org.uk

CONSERVATION PROJECT OFFICER
Carl Allott
Carl’s role allows him to combine a love for conservation with a passion for plants. He is responsible for the conservation breeding project for pine hoverflies and blood-red longhorn beetles, and the implementation of the Biodiversity Action Plan at Highland Wildlife Park.
Contact: callott@rzss.org.uk
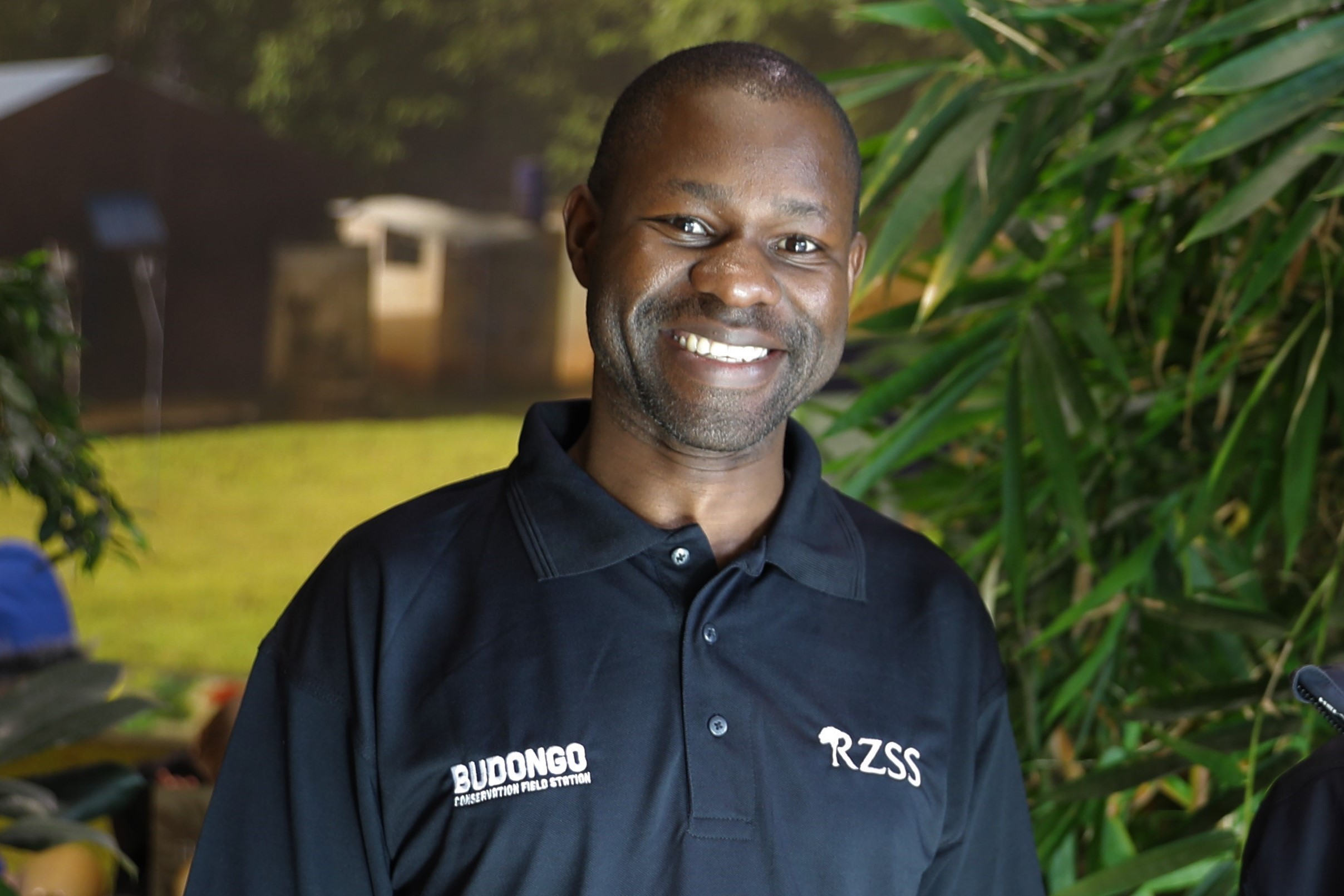

CONSERVATION ASSOCIATE
David Eryenyu
David is the Director of the Budongo Conservation Field Station. He is also currently working towards his PhD on rainforest nutrient cycling.
Contact: eryenyudave@gmail.com
/dr_arnaud_desbiez_checks_armadillo_gala_after_health_check.jpg)
CONSERVATION ASSOCIATE
Dr Arnaud Desbiez
Arnaud Desbiez founded an NGO called ICAS in Brazil (Wild Animal Conservation Institute) to provide administrative support to the two projects he coordinates: the Giant Armadillo Conservation Program and Anteaters and Highways.
Contact: adesbiez@rzss.org.uk
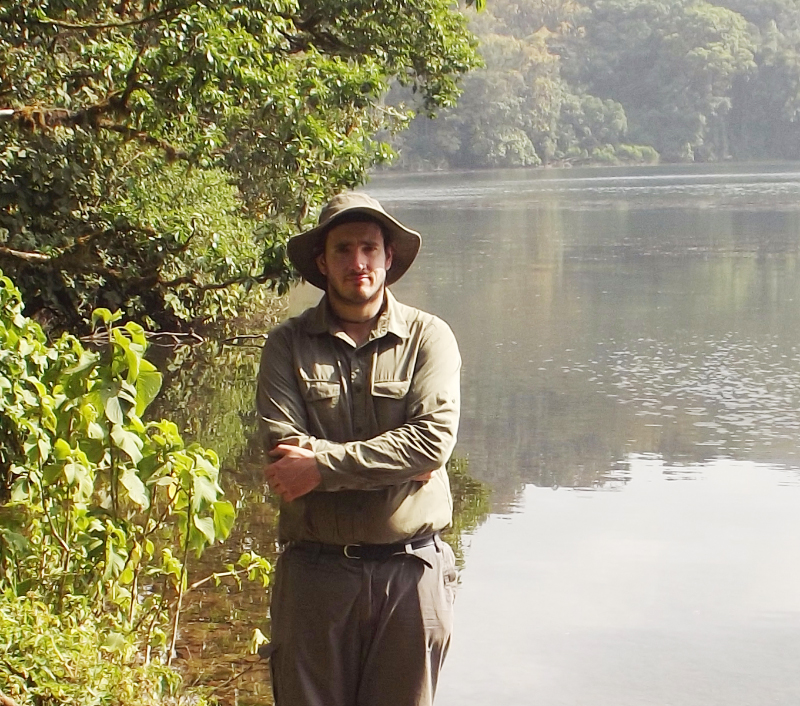

CONSERVATION ASSOCIATE
Dr Thomas Doherty-Bone
Thomas co-ordinates the RZSS African Amphibian & Reptile Program. This project was founded after an RZSS funded expedition to Cameroon in 2006.
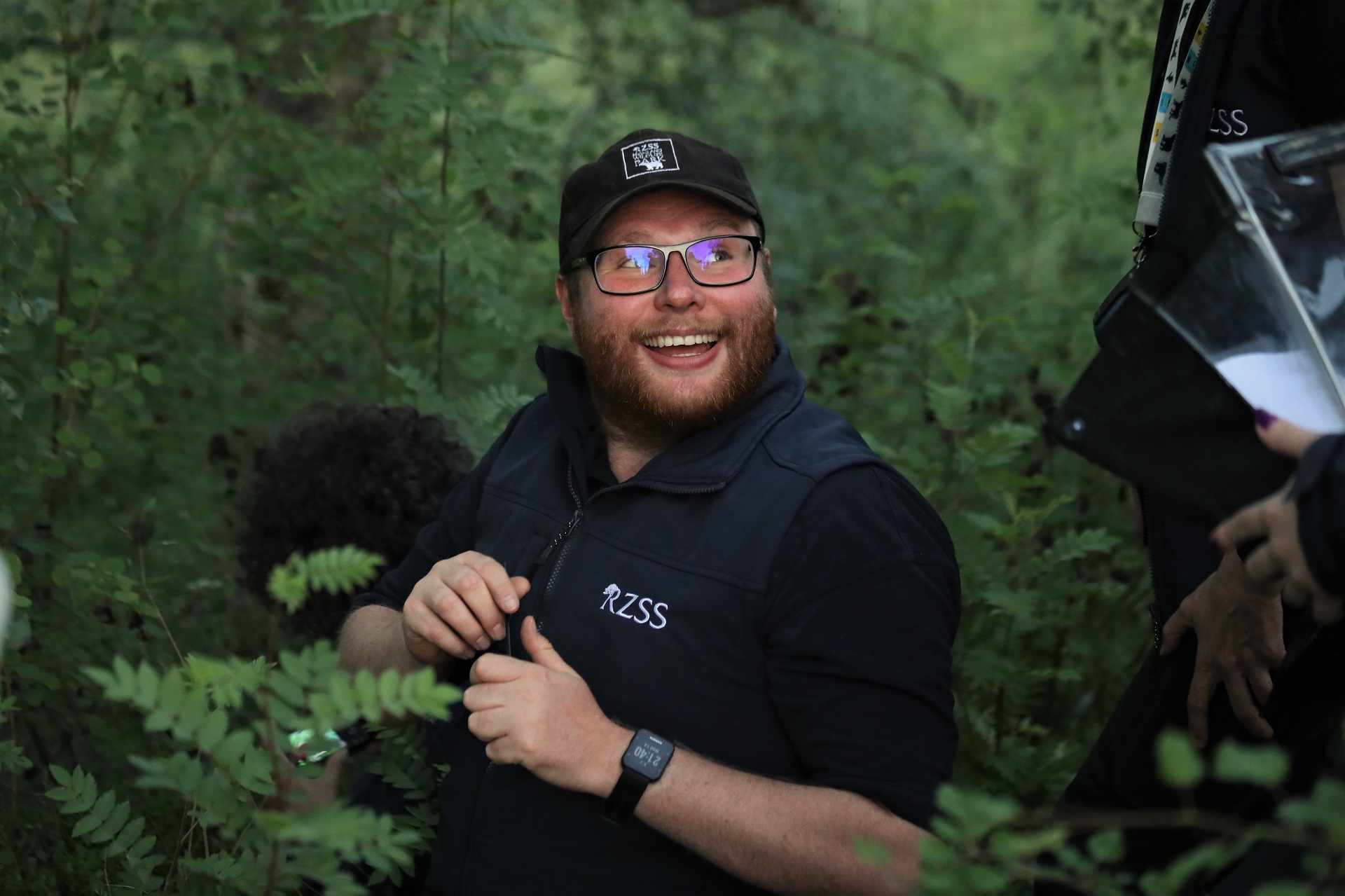

EXPERIENCED ANIMAL KEEPER
Adam Button
Adam works as an Experienced Invertebrate Keeper on the conservation team at Highland Wildlife Park. His focus is on the invertebrate conservation breeding programs with the Dark Bordered Beauty moth and Medicinal leech whilst assisting with the other native invertebrate projects.
Contact: abutton@rzss.org.uk
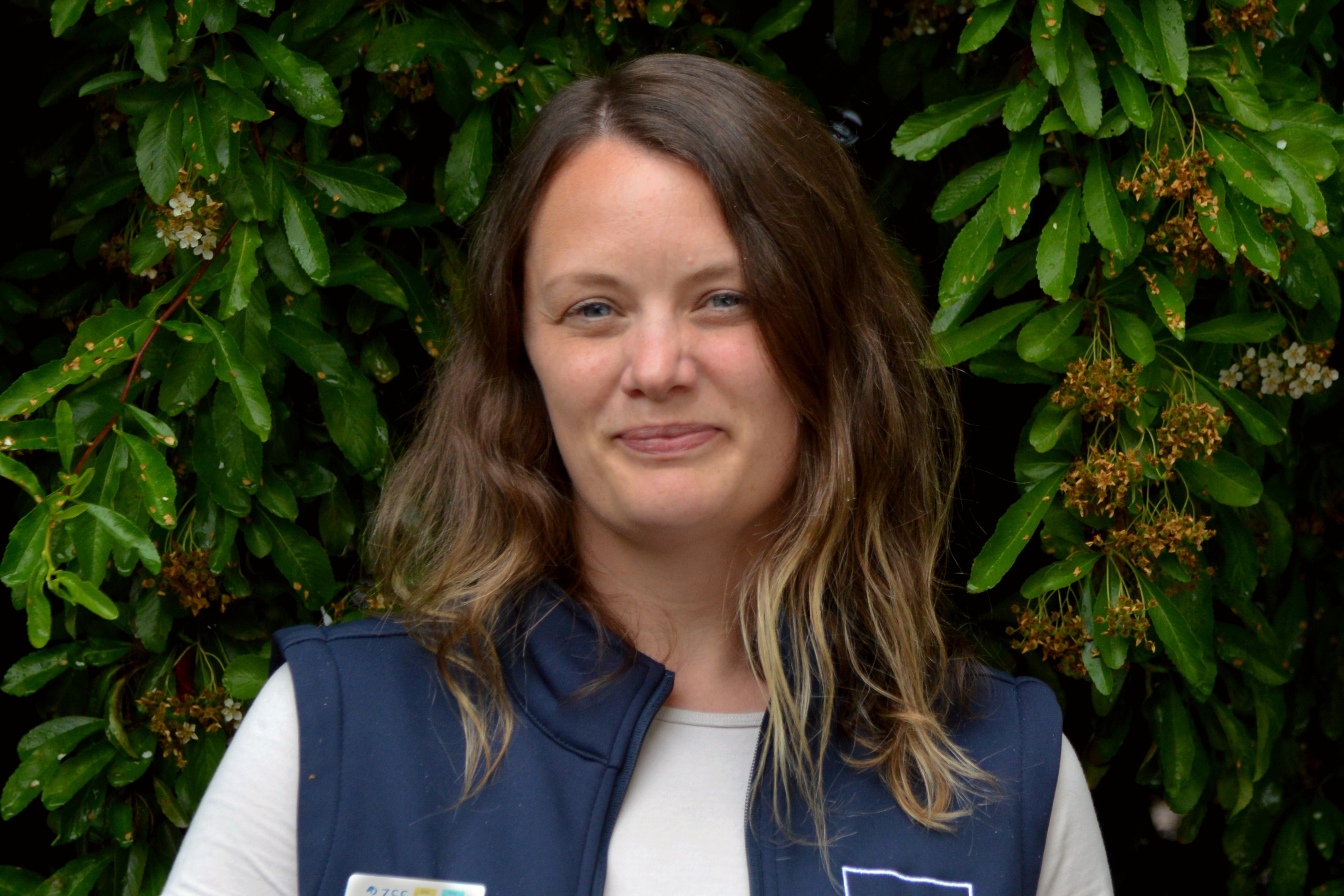

BIOBANK COORDINATOR (RZSS WildGenes)
Dr Cecilia Langhorne
Cecilia is a senior biobank technician in the WildGenes team, overseeing the daily running of the RZSS WildGenes Biobank – a genetic resource holding thousands of biological samples. Cecilia facilitates access to samples by researchers to support global conservation projects through biobank partnerships with the EAZA Biobank and CryoArks network.
Contact: clanghorne@rzss.org.uk

ASSISTANT CONSERVATION PROJECT OFFICER
Laura Daniels
Laura is a conservation scientist working to deliver our projects based in Africa, including leading the Nahan’s partridge project in Uganda. She also works on our pond mud snail project based in Scotland and supports the conservation team, ensuring the department runs smoothly.
Contact: ldaniels@rzss.org.uk

GENOMICS TECHNICIAN (RZSS WildGenes)
Magdalena Butowska
Magda is the genomics technician at RZSS WildGenes, the conservation genetics laboratory at Edinburgh Zoo. She works on generating genomic and genetic data used by the RZSS scientists, contributing to the RZSS conservation projects.
Contact: mbutowska@rzss.org.uk

RESEARCH ASSISTANT (RZSS WildGenes)
Sam Mitchell
Sam is a research assistant in the RZSS WildGenes team with a background in ecology and evolutionary genetics. Sam’s role involves analysis and interpretation of the data generated by the laboratory team, preparing summary reports for conservation partners that will help them make better informed conservation management decisions.
Contact: smitchell@rzss.org.uk

RESEARCH SCIENTIST (RZSS WildGenes)
Dr Jo Howard-McCombe
Jo works as part of the RZSS WildGenes team, processing and analysing genetic data to support a variety of our conservation projects. Currently Jo is focusing on desert antelope and gazelles, giraffes and wildcat projects.
Contact: jhmccombe@rzss.org.uk
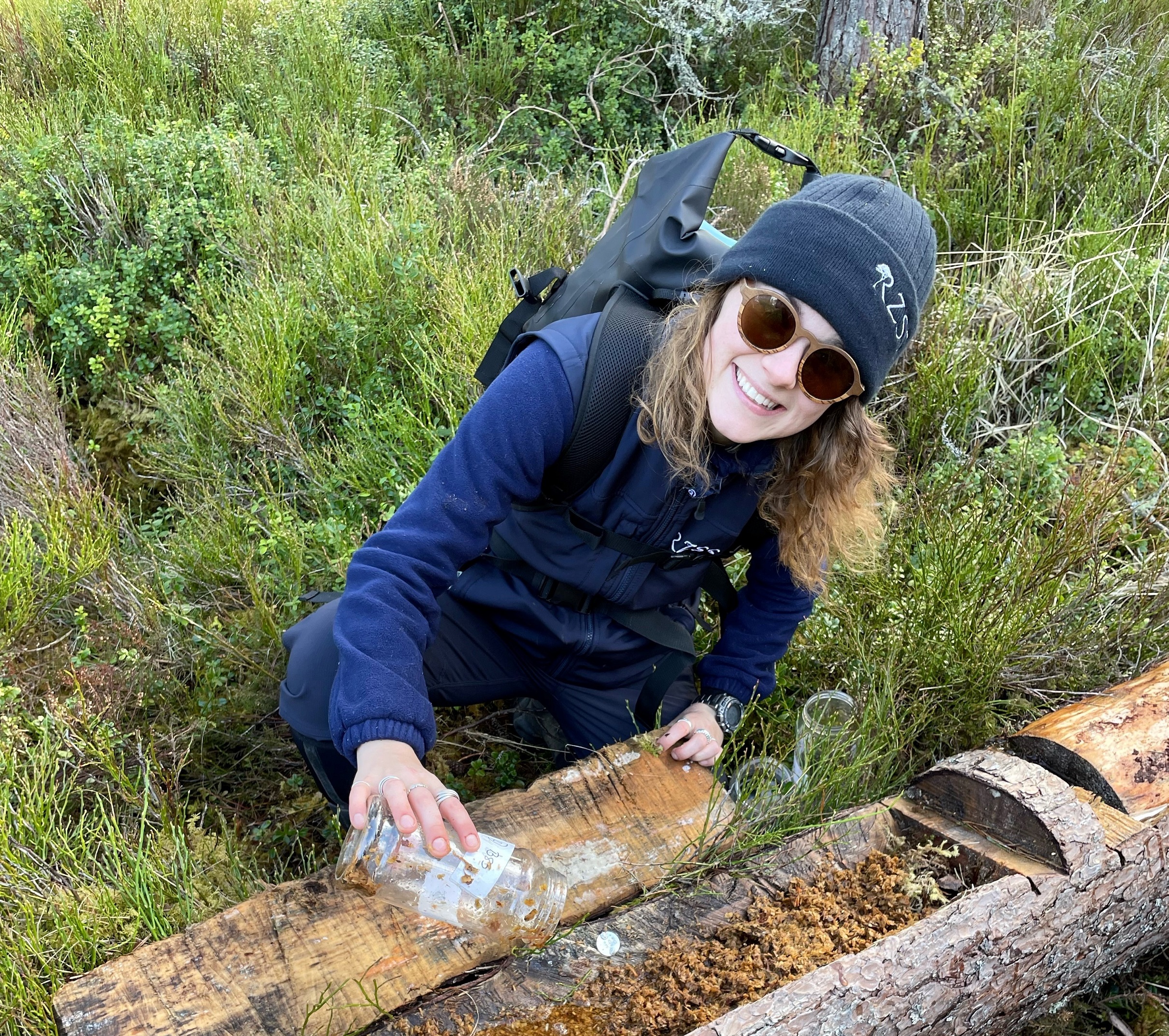

CONSERVATION MANAGER
Georgina Lindsay
Georgina leads the field team at Highland Wildlife Park. She manages various invertebrate conservation breeding and translocation projects including pine hoverflies, dark bordered beauty moths, and medicinal leeches. She is also responsible for overseeing the park's Biodiversity Action Plan.
Contact: glindsay@rzss.org.uk
Saving Wildcats project team

CONSERVATION MANAGER (EX-SITU)
David Barclay
David is the Ex-situ conservation manager and manages our wildcat Conservation Breeding for Release Centre at Highland Wildlife Park. He is also the UK breeding programme coordinator for wildcats, the European and International Breeding programme coordinator for Pallas’s cats and the EAZA Felid TAG co-chair.
Contact: dbarclay@rzss.org.uk

FIELD OPERATIONS MANAGER
Louise Hughes
Louise leads on planning both the long-term and day-to-day activities of the team responsible for pre and post-release monitoring of the wildcats. As well as coordinating field team logistics, surveying an extensive network of camera traps and tracking released wildcats, Louise manages stakeholder engagement objectives and relationships with landowners and land managers.
Contact: lohughes@rzss.org.uk

OUTREACH AND ENGAGEMENT MANAGER
Helena Parsons
Helena is responsible for overseeing the outreach and engagement activities of Saving Wildcats, which includes the project’s community engagement, communications and fundraising. She has a background in felid conservation and public engagement, with an interest in using social science methodologies to engage communities with wildlife restoration.
Contact: hparsons@rzss.org.uk
Veterinary team

HEAD OF VETERINARY SERVICES
Prof Simon Girling
Simon manages the RZSS veterinary team including input for Saving Wildcats and other conservation projects. He set up the UK’s first Residency training programme for the European College of Zoological Medicine (ECZM) in Zoo Health Management in 2013 and is a past President. He is Chair of the UK’s Zoo Experts Committee, member of the UK’s Animal Welfare Committee and the Scottish Animal Welfare Commission.
Contact: sgirling@rzss.org.uk
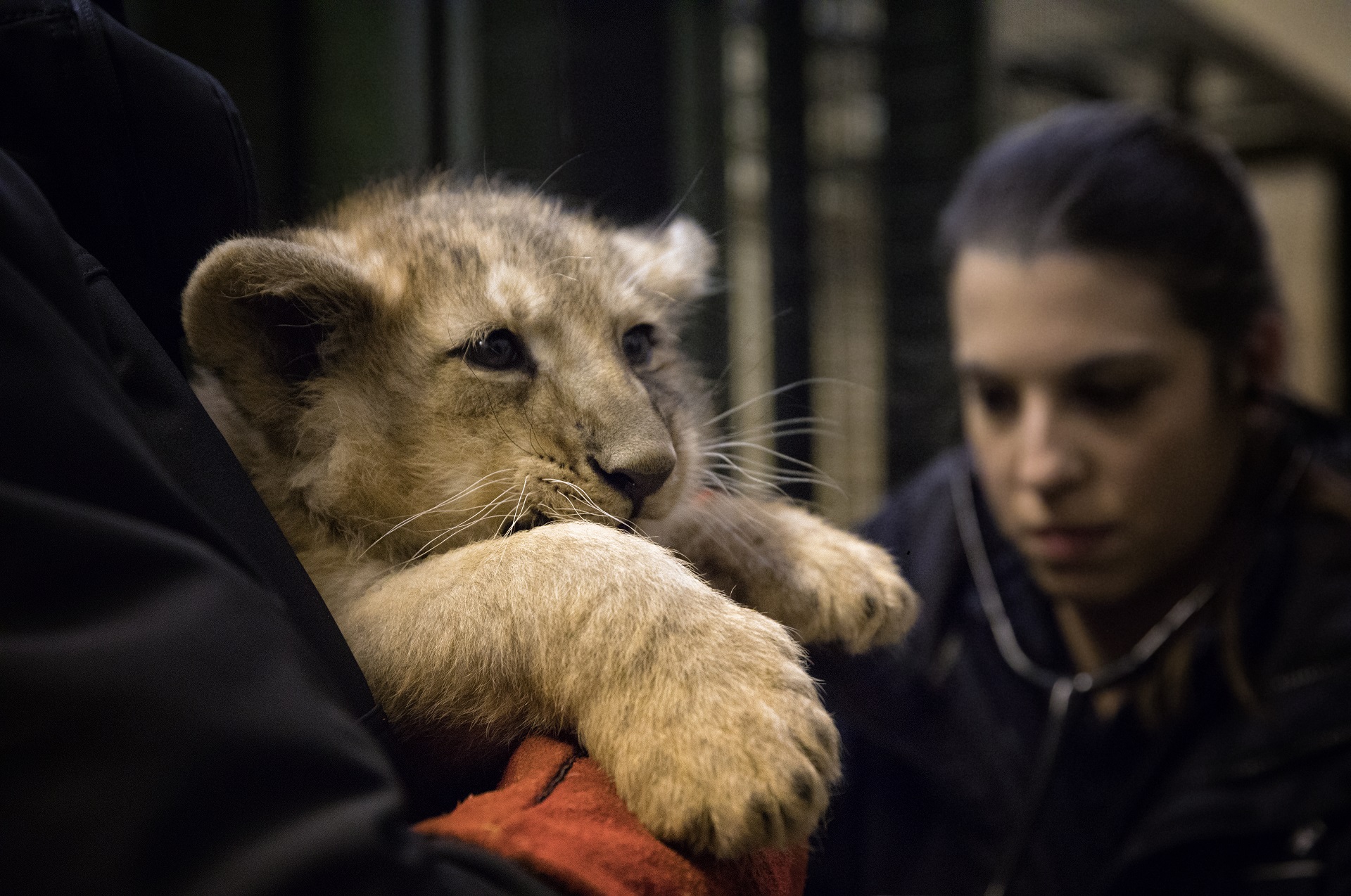

VETERINARY SURGEON
Dr Stephanie Mota
Steph is a veterinary surgeon at the Royal Zoological Society of Scotland, providing veterinary services to the zoo and wildlife conservation projects. She is an EBVS® European Veterinary
Specialist in Zoo Health Management and the EAZA EEP Veterinary Advisor for the Gentoo penguin.
Contact: smota@rzss.org.uk
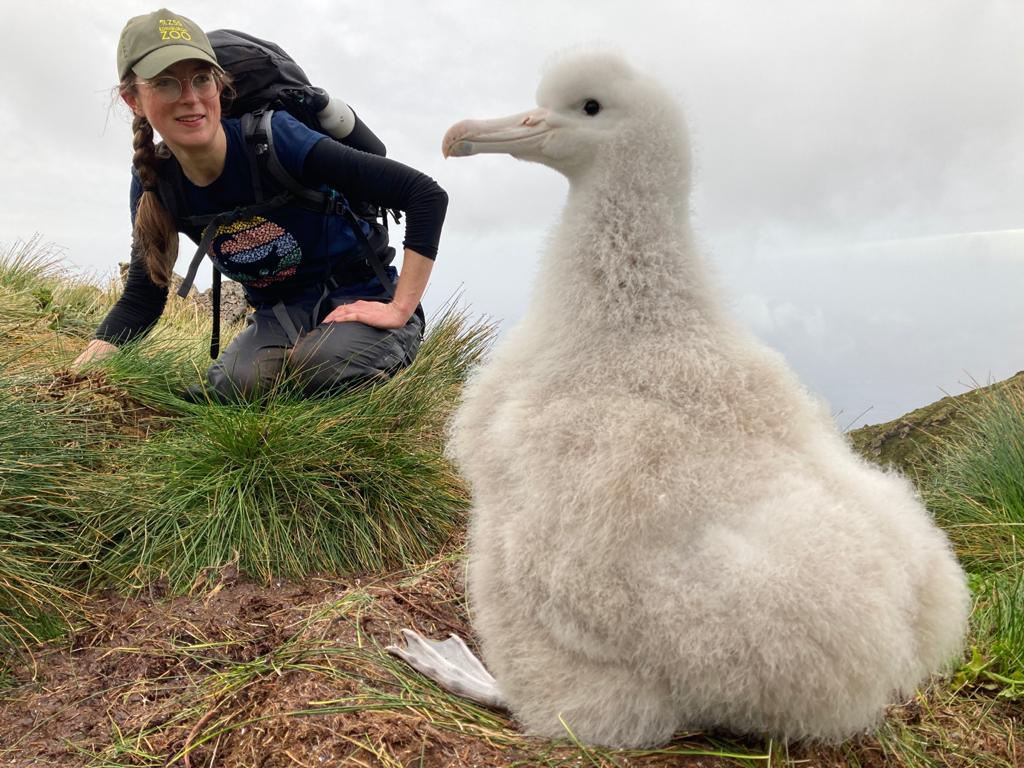

VETERINARY SURGEON
Dr Georgina Cole
Georgina is a zoo and wildlife veterinarian with experience in conservation medicine and animal health research in a range of species. She has a special interest in aquatic animal medicine and is leading on a project investigating aspects of health in the Critically Endangered flapper skate in Scotland.

VETERINARY SURGEON
Dr Alice Bacon

VETERINARY SURGEON
Dr Rebecca Amos
Dr Rebecca Amos is the lead vet for the RZSS ex-situ invertebrate conservation projects. In addition, she assists on the Saving Wildcats conservation project whilst also providing veterinary care for the diverse animal collection at the Highland Wildlife Park.
Contact: ramos@rzss.org.uk

REGISTERED VETERINARY NURSE
Louise Stott
I qualified as a Veterinary Nurse in 2017 after completing the Veterinary Nursing BSc degree. I have completed certificates in Emergency and Critical Care and Advanced Veterinary Nursing Zoo Animal certificate. Louise has worked in first opinion, emergency, referral and teaching before joining the team in May 2022.
Contact: lstott@rzss.org.uk
REGISTERED VETERINARY NURSE
Hannah Jane Brazier Bsc Hons RVN DipVNZS
Hannah is a Registered Veterinary nurse with a diploma in Veterinary nursing of Exotic and zoological species. She works with the wide variety of species at the zoo, from the tiny Partula snails to the giraffes. She works with the Veterinary team to maintain animal health welfare.
Contact: hbrazier@rzss.org.uk

VETERINARY SURGEON
Barbara Ferreira
After working in Southern Africa as a wildlife vet for different conservation projects, she is the most recent member of the RZSS veterinary team with a passion for wild felids.
Contact: bferreira@rzss.org.uk

