How techniques developed by RZSS WildGenes scientists are informing wildcat conservation
24/07/2023 in RZSS
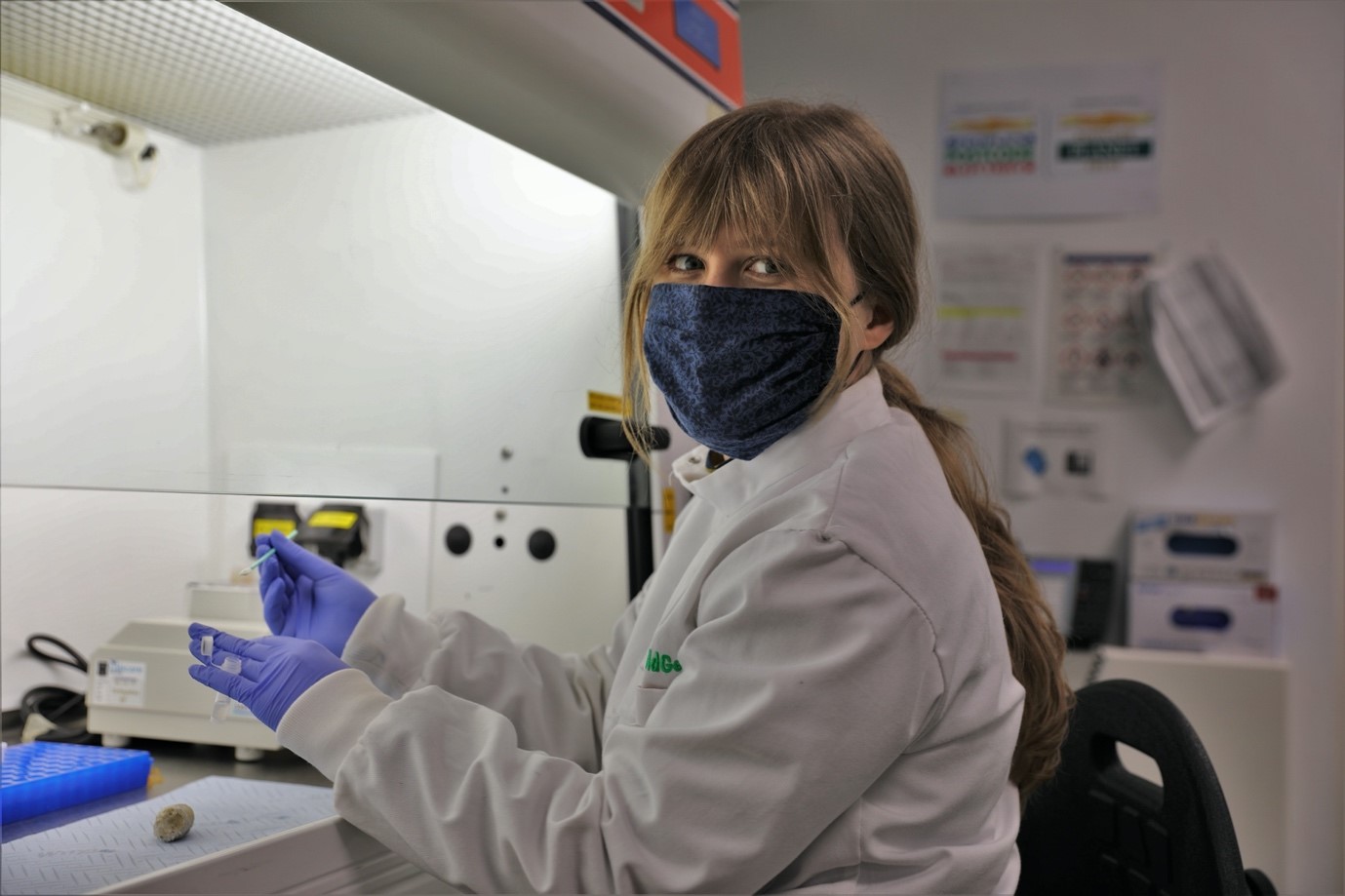

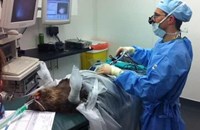
Processing some of the faecal samples from the Highland Wildlife Park at the RZSS WildGenes Lab. Credit: RZSS
My name is Magda Butowska and last year I completed my year-long work placement internship at the RZSS WildGenes Lab, which was a part of my undergraduate master’s programme in Zoology. I am very lucky to have been given the position of a laboratory intern in the lab, where I worked on a variety of projects and species, from pine hoverflies to penguins, gaining invaluable experience.
My main focus during the placement was developing a protocol for analysing the diet of the European wildcats (Felis silvestris) using faecal samples. RZSS leads the Saving Wildcats partnership to restore the species in Scotland, as it is one of Scotland’s most endangered mammals and the last native felid. The project is based at Highland Wildlife Park, where the Saving Wildcats conservation breeding for release centre prepares wildcats for release into the Cairngorms Connect area of the Cairngorms National Park. As the wildcats cannot be fed live prey while in captivity it is vital to use other methods to assess their hunting abilities before and after release.
The European wildcat is a secretive species, and so it is difficult to observe their hunting and feeding behaviour in the wild. Analysing their scats is an alternative way to find out what they eat and therefore how they fit into the ecosystem. Traditional methods involve looking for bones and remains of prey in the scat, which is time-consuming and not always exhaustive in detecting all prey species. Instead, we used a molecular approach, using a technique called DNA metabarcoding, which allowed us to detect the DNA of different prey species in the scat samples. Many samples can be processed together with this technique, and we found, using samples of wildcats with known diets (individuals that are part of the Saving Wildcats project), that we can reliably detect the rabbits, mice, rats, chicken and pheasant that formed their diet.

"Ordie" in the Saving Wildcats conservation breeding for release centre. Credit: Saving Wildcats
The protocol we developed will be a useful tool for monitoring the feeding, diet components and hunting ability of wildcats. It can also be easily adjusted for other feline species’ and forms part of a wider project that uses genetics to analyse carnivore diets. I had the opportunity to present these findings as a poster at the European Conservation Genetics conference (ConsGen22), held in Edinburgh last September.
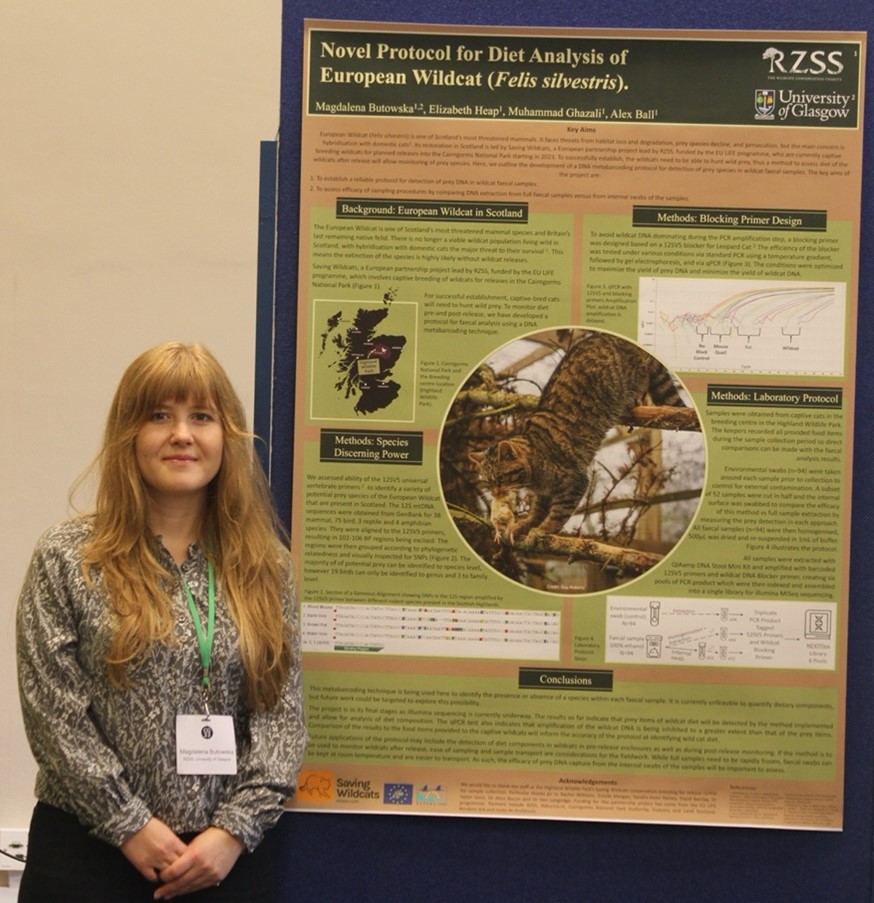

Presenting the results at the ConsGen22 Conference in Edinburgh. Credit: RZSS
From Saturday 22 July to Sunday 30 July, get involved with Poo Fest at Edinburgh Zoo. Find out more at edinburghzoo.org.uk/events/poo-fest/.
If you would like to learn more about the Saving Wildcats partnership you can visit the project website at savingwildcats.org.uk.
To learn more about RZSS WildGenes, explore our conservation genetics projects.
Featured Articles

An update from the Budongo Forest
19/04/2024 in Conservation

Edinburgh Zoo named best zoo in Scotland
15/04/2024 in Edinburgh Zoo























Follow EZ